Published papers




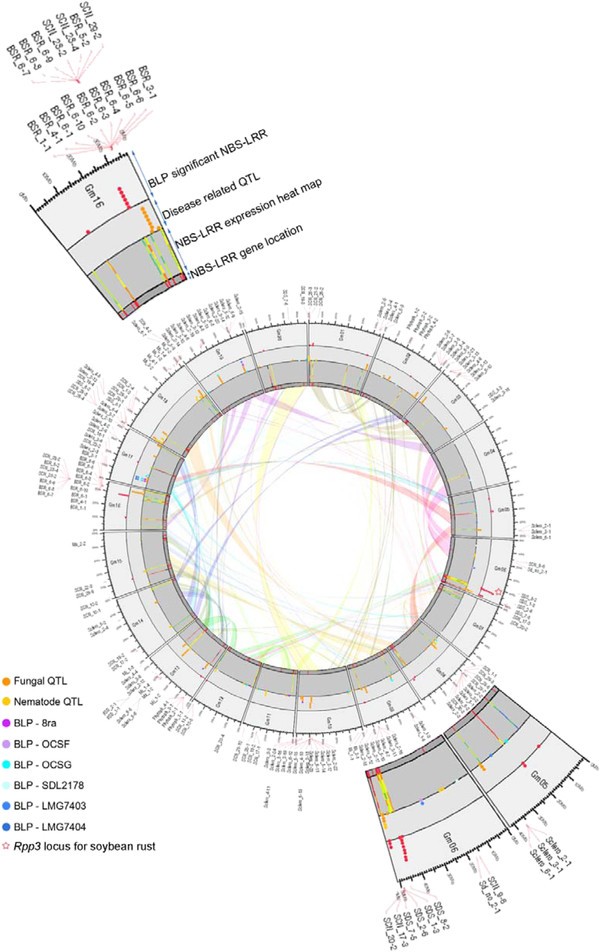
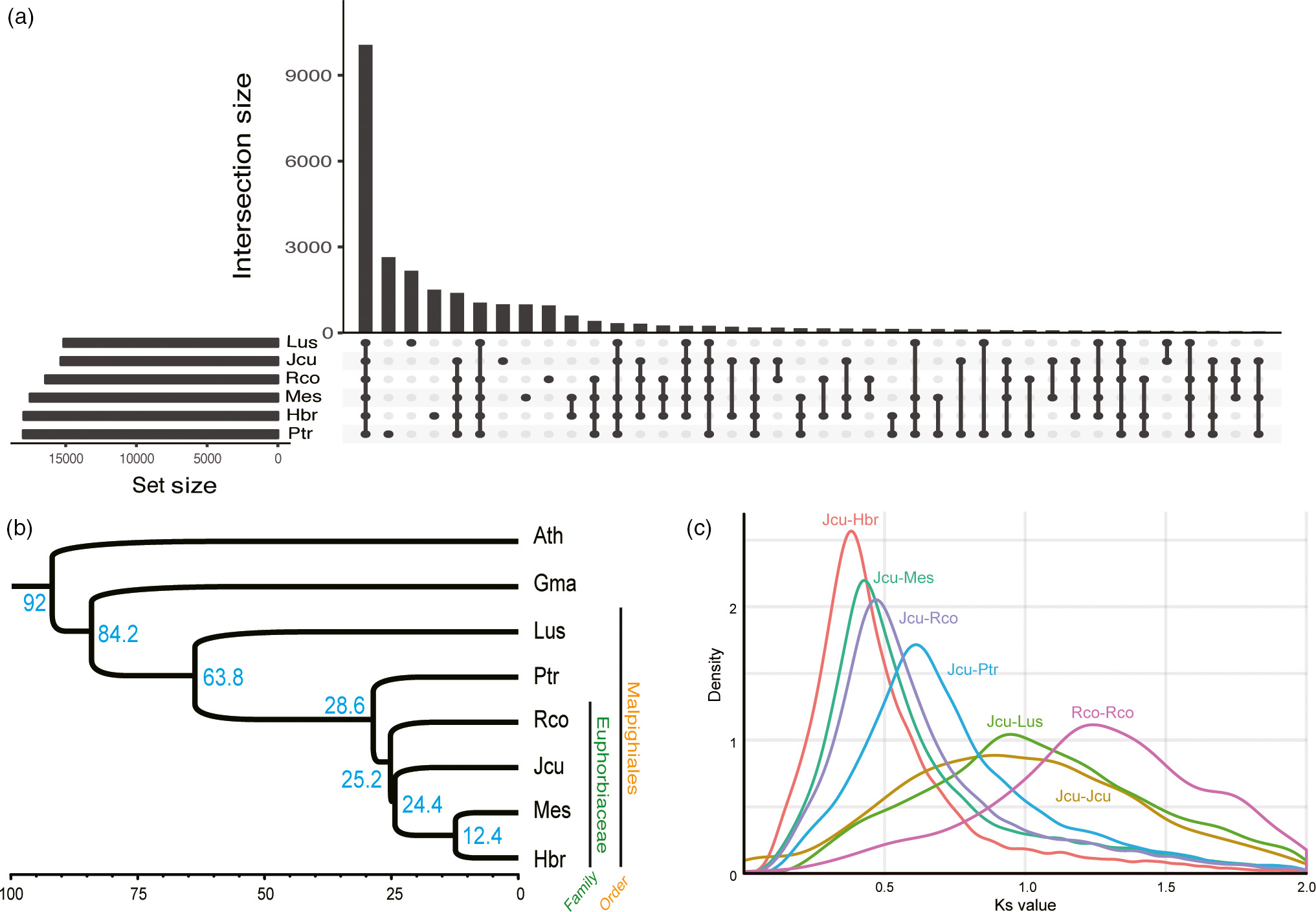
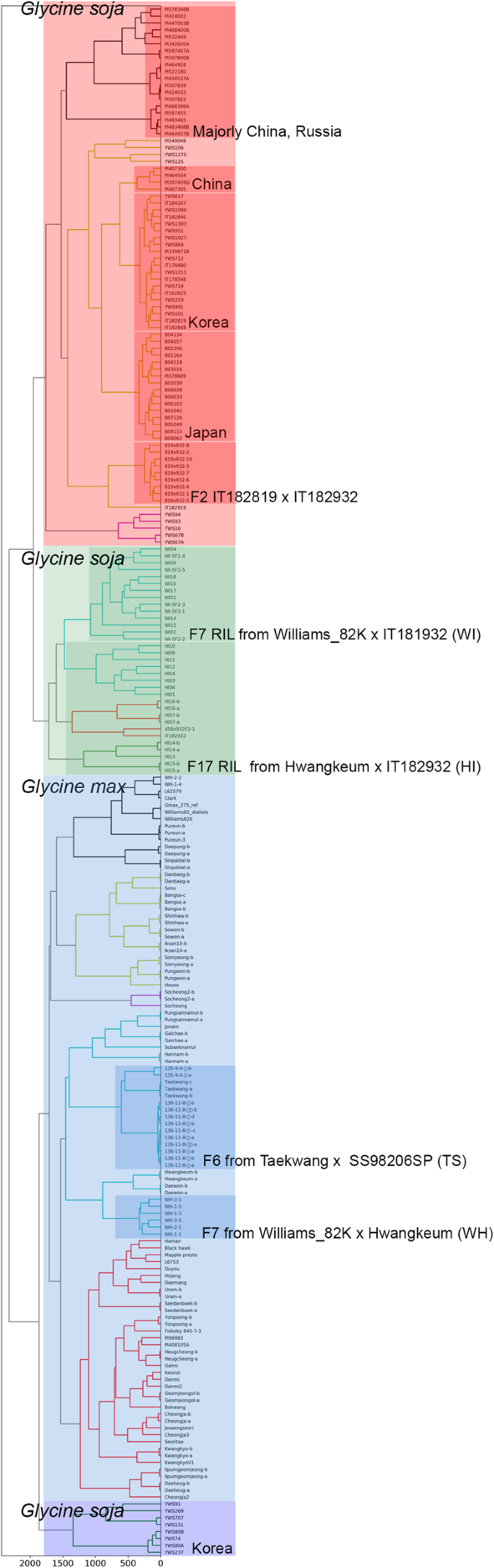


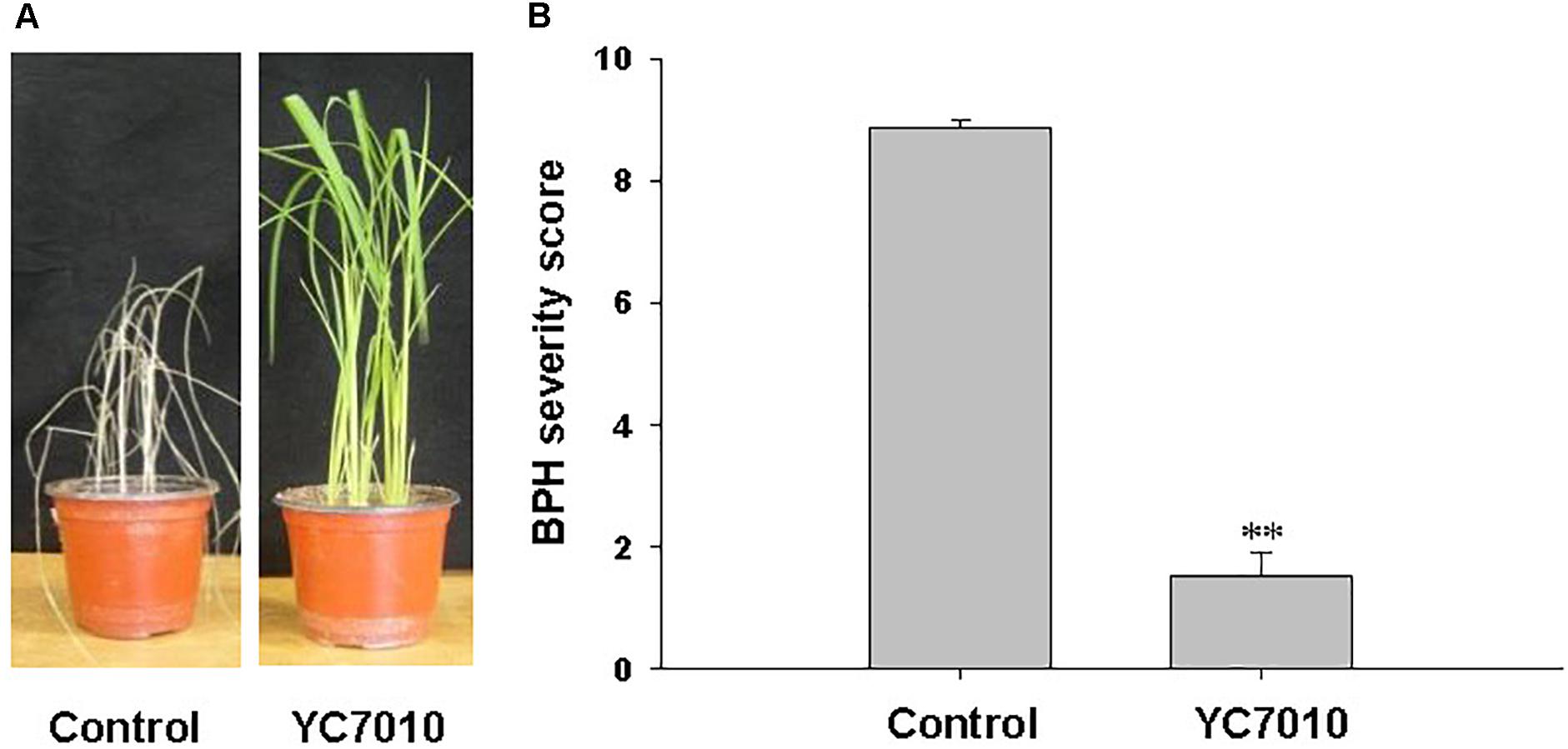
생물학은 생물현상을 관찰하고 그에 맞는 가설을 세우고 꾸준히 증명해나가는 학문입니다. 생물현상의 관찰값이 많지 않던 과거와는 달리 최근 기술의 발전에 따라 생물 빅 데이터라고 불리울 만큼 많은 데이터를 생산하고 있습니다. 이에 생물현상의 관찰값이 대량화되어 “생물정보”를 다루는 학문이 정착하였는데 그것이 생물 정보학입니다. 특히 저렴한 가격으로 꾸준히 생산되고 있는 생물정보는 Next generation sequencer (NGS) 데이터입니다. Genome,Transcriptome, Epigenome, Interactome 등의 데이터가 과거에 비해 빠른속도로 축적되고 있는데 이를 통합하여 생명현상을 충분히 설명할 수 있는 모델을 만들고자 하는 것이 본 실험실의 주 목적입니다. 이는 생물 유전자원을 쉽게 대표할 수 있는 1. Genome profile 개발 그리고 2. Genome profile 기반의 인공지능 예측 모델을 통한 작물육종 시스템 개발 3, Portable sequencer를 통한 유전체 기반 검역시스템을 개발하여 실용화 하기위해 노력하고 있습니다. 앞으로 생물정보학은 생물학을 수행하기 위해 필수 학문이 될것으로 예상됩니다. 끊임없이 쏟아져 나오는 데이터들은 우리가 이미 알고 있던 가설들을 무너뜨리기도 하고 더욱더 강력한 가설들을 제시할 것 입니다. 또한 생물학이라는 분야에서 이루어낸 성취들을 실생활에 적용하는데 역시 생물정보학은 중요한 역할을 할 것으로 기대합니다. 이러한 생물정보시대에 대비한 잘 훈련된 젊은 연구자들이 미래의 생물학을 이끌어나갈 것이라고 생각합니다.









